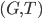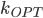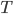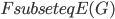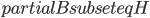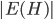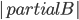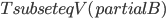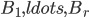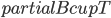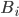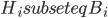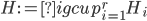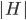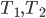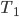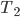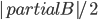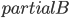The Graph Isomorphism problem is interesting for many reasons, one being the fact that its complexity status is still unclear - it is probably not NP-complete, but not known to be in P either. In the past decades, researchers identified a large number of special graph classes for which GI is polynomial-time solvable, including planar graphs, graphs of bounded maximum degree, and graphs excluding a fixed minor, among others.
A closer look at the known algorithms for graphs of bounded degree or graphs excluding a fixed minor reveals that the dependency in the running time bound on the "forbidden pattern" is quite bad: 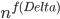 in the first case or
in the first case or 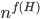 in the second case, for some function
in the second case, for some function 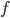 . Seeing such dependencies, a natural question is: do there exist fixed-parameter algorithms, parameterized by the "forbidden pattern"? In other words, does the exponent in the polynomial factor in
. Seeing such dependencies, a natural question is: do there exist fixed-parameter algorithms, parameterized by the "forbidden pattern"? In other words, does the exponent in the polynomial factor in 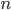 need to depend on the pattern, on can we obtain an algorithm with running time
need to depend on the pattern, on can we obtain an algorithm with running time 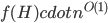 ?
?
This question remains widely open for all aforementioned cases of bounded degree or excluded minor. In our recent FOCS 2014 paper, we have answered the question affirmatively for the special case of bounded treewidth graphs, proving that isomorphism of two 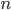 -vertex graphs of treewidth
-vertex graphs of treewidth 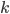 can be tested in time
can be tested in time 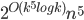 .
.
As a starting point, observe that comparing (testing isomorphism) of two graphs together with their given tree decompositions (that is, we want to check if the whole structures consisting of a graph and its decomposition are isomorphic) can be done easily in time polynomial in the size of the graphs, and exponential in the width of the decompositions. This is a relatively standard, but tedious, exercise from the area of designing dynamic programming algorithms on tree decompositions: if you have seen a few of these, I guess you can figure out the details, but if you don't, this is probably one of the worst examples to start with, so just assume it can be done. Alternatively, you may use the recent results of Otachi and Schweitzer that say that one needs only a set of bags for the decomposition, even without the structure of the decomposition tree itself.
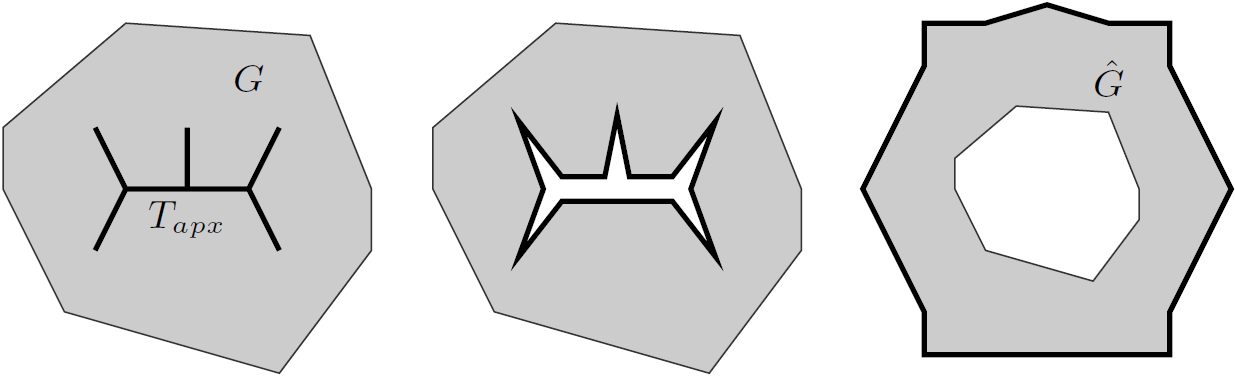
With this observation, the task of comparing two bounded-treewidth graphs reduces to the following: given a graph 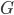 of treewidth
of treewidth 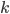 , we would like to compute in an isomorphic-invariant way a tree decomposition of $G$ of near-optimal width. Here, isomorphic-invariant means that we do not want to make any decisions depending on the representation of the graph in the memory (like, "take arbitrary vertex"), and the output decomposition should be the same (isomorphic) for isomorphic input graphs. Given such an isomorphic-invariant treewidth approximation algorithm, we can now compute the decompositions of both input graphs, and compare given pairs (graph, its tree decomposition), as described in the previous paragraph. Note that for this approach we do not need to output exactly one tree decomposition; we can output, say,
, we would like to compute in an isomorphic-invariant way a tree decomposition of $G$ of near-optimal width. Here, isomorphic-invariant means that we do not want to make any decisions depending on the representation of the graph in the memory (like, "take arbitrary vertex"), and the output decomposition should be the same (isomorphic) for isomorphic input graphs. Given such an isomorphic-invariant treewidth approximation algorithm, we can now compute the decompositions of both input graphs, and compare given pairs (graph, its tree decomposition), as described in the previous paragraph. Note that for this approach we do not need to output exactly one tree decomposition; we can output, say, 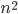 of them, and compare every pair of decompositions for two input graphs for the Graph Isomorphism problem; we only need that the set of output decompositions is isomorphic invariant.
of them, and compare every pair of decompositions for two input graphs for the Graph Isomorphism problem; we only need that the set of output decompositions is isomorphic invariant.
To cope with this new task, let us look at the known treewidth approximation algorithms; in particular, the arguably simplest one - the Robertson-Seymour algorithm. This algorithm provides a constant approximation in 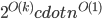 time, where
time, where 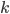 is the treewidth of the input
is the treewidth of the input 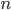 -vertex graph
-vertex graph 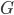 . It is a recursive procedure that decomposes a part
. It is a recursive procedure that decomposes a part 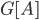 of the input graph, given a requirement that a set
of the input graph, given a requirement that a set 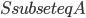 should be contained in the top bag of the output decomposition. Think of
should be contained in the top bag of the output decomposition. Think of 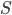 as an interface to the rest of the graph; we will always have
as an interface to the rest of the graph; we will always have 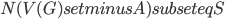 , but for technical reasons the set
, but for technical reasons the set 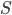 could be larger. During the course of the algorithm we maintain an invariant
could be larger. During the course of the algorithm we maintain an invariant 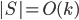 .
.
In the leaves of the recursion we have 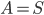 , and we output a single bag
, and we output a single bag 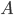 . If
. If 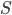 is small, say,
is small, say, 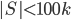 , then we can add an arbitrary vertex to
, then we can add an arbitrary vertex to 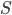 and recurse. The interesting things happen when
and recurse. The interesting things happen when 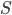 grows beyond the threshold
grows beyond the threshold 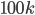 .
.
As the graph is of treewidth 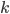 , but
, but 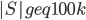 , in an optimal decomposition of
, in an optimal decomposition of 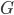 the set
the set 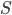 is spread across many bags. A simple argument shows that there exists a bag
is spread across many bags. A simple argument shows that there exists a bag 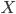 ,
, 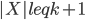 , such that every component of
, such that every component of 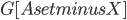 contains at most
contains at most 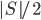 vertices of
vertices of 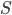 . The algorithm takes
. The algorithm takes 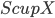 as a root bag (note that
as a root bag (note that 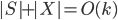 ), and recurses on every connected component of
), and recurses on every connected component of 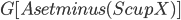 ; formally, for every connected component
; formally, for every connected component 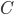 of
of 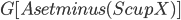 , we recurse on
, we recurse on 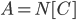 with
with 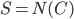 . Since every component of
. Since every component of 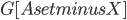 contains at most
contains at most 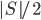 vertices of
vertices of 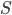 , in every recursive call the new set
, in every recursive call the new set 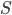 is of size at most
is of size at most 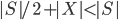 .
.
In the above algorithm, there are two steps that are very deeply not isomorphic-invariant: "take an arbitrary vertex to 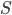 " in the case of small
" in the case of small 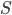 , and "take any such
, and "take any such 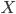 " in the case of large
" in the case of large 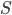 . We start fixing the algorithm in the second, more interesting case.
. We start fixing the algorithm in the second, more interesting case.
Before we start, let us think how we can find such a set 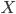 - the argument provided two paragraphs ago only asserted its existence. One of the ways to do it is to iterate through all disjoint sets
- the argument provided two paragraphs ago only asserted its existence. One of the ways to do it is to iterate through all disjoint sets 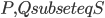 of size exactly
of size exactly 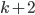 each, and check if the minimum separator between
each, and check if the minimum separator between 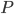 and
and 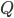 in
in 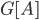 is of size at most
is of size at most 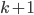 (such a separator can contain vertices of
(such a separator can contain vertices of 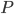 or
or 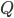 , these are deletable as well). If such a separator is found, it is a good candidate for the set
, these are deletable as well). If such a separator is found, it is a good candidate for the set 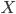 - it does not necessarily have the property that every connected component of
- it does not necessarily have the property that every connected component of 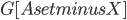 contains at most half of the vertices of
contains at most half of the vertices of 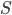 , but the claim that in the recursive calls the size of the set
, but the claim that in the recursive calls the size of the set 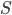 decreased is still true. Of course, in this approach we have two not isomorphic-invariant choices: the choice of
decreased is still true. Of course, in this approach we have two not isomorphic-invariant choices: the choice of 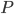 and
and 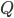 , and the choice of the minimum separator
, and the choice of the minimum separator 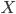 .
.
Luckily, an (almost) isomorphic-invariant way to make the second choice has already been well understood: by submodularity of cuts, there exists a well-defined notion of the minimum cut closest to 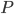 , and the one closest to
, and the one closest to 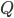 . But how about the choice of
. But how about the choice of 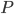 and
and 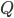 ?
?
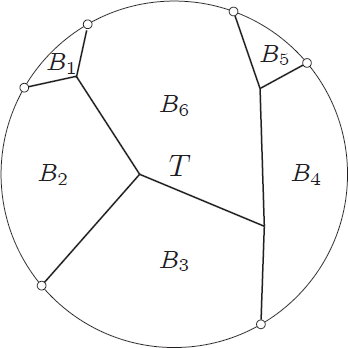
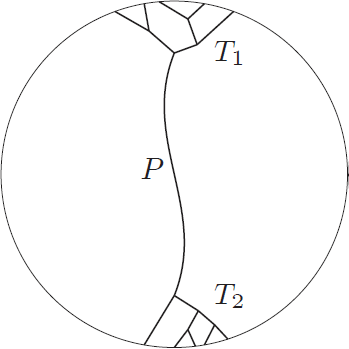
The surprising answer is: it works just fine if we just throw into the constructed bag all separators for every choice of 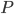 and
and 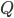 . That is, we start with the tentative bag
. That is, we start with the tentative bag 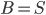 , we iterate over all such (ordered) pairs
, we iterate over all such (ordered) pairs 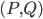 , and throw into the constructed bag
, and throw into the constructed bag 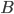 the minimum cut between
the minimum cut between 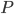 and
and 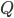 that is closest to
that is closest to 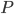 , as long as it has size at most
, as long as it has size at most 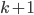 . We recurse on all connected components of
. We recurse on all connected components of 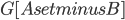 as before; the surprising fact is that the sizes of the sets
as before; the surprising fact is that the sizes of the sets 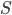 in the recursive calls did not grow. To prove this fact, we analyse what happens to the components of
in the recursive calls did not grow. To prove this fact, we analyse what happens to the components of 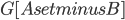 when you add
when you add 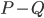 cuts one-by-one; a single step of this induction turns out to be an elementary application of submodularity. Furthermore, note that if the size of
cuts one-by-one; a single step of this induction turns out to be an elementary application of submodularity. Furthermore, note that if the size of 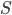 is bounded in terms of
is bounded in terms of 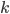 , so is the set of the bag
, so is the set of the bag 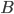 :
: 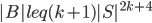 .
.
Having made isomorphic-invariant the "more interesting" case of the Robertson-Seymour approximation algorithm, let us briefly discuss the case of the small set 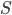 : we need to develop an isomorphic-invariant way to grow this set. Here, we need some preprocessing: by known tricks, we can assume that (a) the input graph does not contain any clique separators, and (b) if
: we need to develop an isomorphic-invariant way to grow this set. Here, we need some preprocessing: by known tricks, we can assume that (a) the input graph does not contain any clique separators, and (b) if 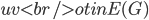 , then the minimum cut between
, then the minimum cut between 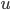 and
and 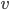 is of size at most
is of size at most 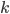 . Furthermore, observe that in our algorithm, contrary to the original Robertson-Seymour algorithm, we always maintain the invariant that
. Furthermore, observe that in our algorithm, contrary to the original Robertson-Seymour algorithm, we always maintain the invariant that  and
and 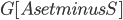 is connected. Thus,
is connected. Thus, 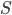 is never a clique in
is never a clique in 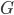 , and, if
, and, if 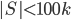 , we can throw into a bag the minimum cut between
, we can throw into a bag the minimum cut between 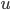 and
and 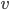 that is closest to
that is closest to 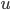 , for every ordered pair
, for every ordered pair 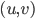 where
where 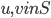 ,
, 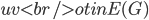 .
.
In this way, the lack of clique separators ensures that we always make a progress in the algorithm, by "eating" at least one vertex of 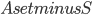 . The bound on the size of minimum cut between two nonadjacent vertices ensures that the bag created in this way is of size
. The bound on the size of minimum cut between two nonadjacent vertices ensures that the bag created in this way is of size 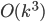 , and so are the sets
, and so are the sets 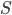 in the recursive calls. Consequently, our algorithm computes in an isomorphic-invariant way a tree decomposition with adhesions of size
in the recursive calls. Consequently, our algorithm computes in an isomorphic-invariant way a tree decomposition with adhesions of size 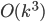 and bags of size
and bags of size 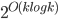 .
.
Well, there is one small catch - neither of the aforementioned steps is good to start the algorithm, to provide the initial call of the recursion; the Robertson-Seymour approximation starts with just 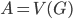 and
and 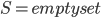 . Here, we can start with
. Here, we can start with 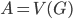 and
and  for every pair
for every pair 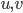 of nonadjacent vertices, as in this call the step with the small set
of nonadjacent vertices, as in this call the step with the small set 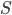 will work fine. But this produces not a single isomorphic-invariant tree decomposition, but a family of
will work fine. But this produces not a single isomorphic-invariant tree decomposition, but a family of 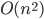 decompositions - which is still fine for the isomorphism test.
decompositions - which is still fine for the isomorphism test.
A cautious reader would also notice that the described algorithm results in a double-exponential dependency on the parameter 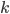 in the running time, contrary to the claim of the
in the running time, contrary to the claim of the 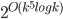 term. Getting down to this dependency requires a bit more technical work; we refer to the full version of our paper for details.
term. Getting down to this dependency requires a bit more technical work; we refer to the full version of our paper for details.